- Software
- Open access
- Published:
Sealer: a scalable gap-closing application for finishing draft genomes
BMC Bioinformatics volume 16, Article number: 230 (2015)
Abstract
Background
While next-generation sequencing technologies have made sequencing genomes faster and more affordable, deciphering the complete genome sequence of an organism remains a significant bioinformatics challenge, especially for large genomes. Low sequence coverage, repetitive elements and short read length make de novo genome assembly difficult, often resulting in sequence and/or fragment “gaps” – uncharacterized nucleotide (N) stretches of unknown or estimated lengths. Some of these gaps can be closed by re-processing latent information in the raw reads. Even though there are several tools for closing gaps, they do not easily scale up to processing billion base pair genomes.
Results
Here we describe Sealer, a tool designed to close gaps within assembly scaffolds by navigating de Bruijn graphs represented by space-efficient Bloom filter data structures. We demonstrate how it scales to successfully close 50.8 % and 13.8 % of gaps in human (3 Gbp) and white spruce (20 Gbp) draft assemblies in under 30 and 27 h, respectively – a feat that is not possible with other leading tools with the breadth of data used in our study.
Conclusion
Sealer is an automated finishing application that uses the succinct Bloom filter representation of a de Bruijn graph to close gaps in draft assemblies, including that of very large genomes. We expect Sealer to have broad utility for finishing genomes across the tree of life, from bacterial genomes to large plant genomes and beyond. Sealer is available for download at https://github.com/bcgsc/abyss/tree/sealer-release.
Background
De novo assembly using next-generation sequencing short read sequences have been successful in producing draft genome sequences [1]. However, complete and fully automated assembly of genomes remains elusive, especially for prohibitively sized genomes such as human. Problems generally reside in areas of low-coverage or highly repetitive sequences. Even in cases where the overall long-range sequence structure can be disambiguated, on shorter scales there may be ambiguous or undetermined bases, producing regions of Ns or “gaps” in assembly scaffolds. The need for gap closure is made more evident for large (in giga-base pair, Gbp, range) genomes, such as in H. sapiens, where there are higher occurrences of complex genomic features. In fact, as the cost of DNA sequencing decreases faster than the cost of computer hardware, more raw sequencing data will be generated while computational resources will remain mostly the same [2]. Thus it is critical to develop tools that can scale up to these large datasets while using minimal computing resources. Projects such as the 1000 Genomes Project [3], The Cancer Genome Atlas [http://cancergenome.nih.gov/], and clinical uses of whole-genome sequencing [4] highlight the trend of processing Gbp-scale datasets.
Even though these projects are about re-sequencing human genomes and transcriptomes, it was demonstrated that de novo assembly of the raw reads provides valuable information on structural variations [5–8]. Thus these initiatives would benefit from a gap-closing tool that can improve the quality of human draft assemblies, while having low runtime and memory usage as it would help reduce the cost of analysis [4].
Obviously, gap-closing algorithms are also valuable in de novo sequencing projects, with some of the contemporary studies using the concept to improve assembly contiguity [9]. When the closed gaps refine the sequence content in or near genic or regulatory regions, they provide information for the downstream annotation work, and enable biological insights.
Hence, several tools have been designed to close gapped regions with sequence reads, including BaseClear GapFiller [10] and SOAPdenovo GapCloser [11]. The former implements a method that seeks read pairs with one pair aligning within a contig and its mate partially located in a region identified as a gap. These partially aligned reads are used to close the gap through sequence overlap. With GapCloser, a stand-alone tool in the SOAPdenovo package, reads are aligned to contig positions, and a base extension algorithm is performed. Although both of these tools have been shown to successfully close gaps in Mega-bp scale datasets such as in S. cerevisiae (11 Mbp) and human chromosome 14 (95 Mbp) genomes, they have difficulties to process larger datasets, such as the entire H. sapiens genome.
To address this need, we developed Sealer, a resource-efficient gap-filling software. Sealer uses an assembly utility within the ABySS package, called Konnector [12] as its engine to close intra-scaffold gaps. We demonstrate the scalability of Sealer on the white spruce (P. glauca) draft genome [13], which it processes under 27 h using 40 GB RAM – resources that can be found in contemporary commodity desktop computers. We evaluate Sealer by running tests on experimental datasets, comparing runtime, memory usage, and gap-closing success rate against state-of-the-art gap-filling applications. We expect Sealer to find a wide application for finishing small and large genomes alike.
Implementation
Algorithm Overview
Sealer performs three sequential functions (Additional file 1: Figure S1). First, regions with Ns are identified from an input scaffold file, and nucleotides flanking each gap are extracted. Then, flanking sequence pairs are used as input to Konnector along with a set of reads with a high level of coverage redundancy. Typically, the reads represent the original dataset from which the draft assembly is generated, or may be reads from further whole genome shotgun (WGS) sequencing data generated from the same sample. The Konnector utility is run with a range of k-mer lengths to connect the flanking sequences. Finally, successfully connected sequences are inserted into the gaps of the original scaffolds, and a new gap-filled scaffold file is generated. Sealer ignores size discrepancies between gaps and newly introduced sequences, since gap sizes are often estimated from fragment library distributions and assemblers do not generally provide confidence intervals for every region of Ns. Despite this, large expansions of the assembly are unlikely due to decreasing gap-closing yield of Konnector as the gap size increases [12]. Below are further details on these three steps.
-
Step 1: Extracting sequences flanking gaps. Sealer identifies regions of Ns in an input assembly. It then searches for flanking sequences that do not contain Ns, and are at least -L nucleotides long (default = 100). The start position and scaffold ID of each gap are recorded for downstream processing.
-
Step 2: Konnector assembly. The underlying engine for Sealer is Konnector, a tool to generate pseudo long reads from paired-end sequencing data by filling the unknown sequence between read pairs using the redundancy in sequence coverage. Given a k-mer length and a read set, Konnector builds a Bloom filters to represent all k-mers in the reads, and retains k-mers that are observed with a certain threshold multiplicity or higher (default = 2). It uses this data structure as an implicit de Bruijn graph [14] to perform a depth-limited, bidirectional, breadth-first graph search for a path that connects the flanking reads. In Sealer, gap-flanking sequence pairs are used in lieu of read pairs.
Sealer invokes Konnector with a range of k-mer lengths. The advantage of this “k sweep” strategy is, gaps with low coverage have an increased chance of being closed by shorter k-mer lengths, while gaps with highly repetitive sequences are more likely to be closed by larger k-mer values. At each k-mer instance, all possible traversals within the depth limit are identified between each flanking sequence pairs. Unique traversals, and traversals with path multiplicity less than or equal to a user-specified threshold (default = 2) are reported as successful connections. Consensus sequences are produced for multiple paths, reporting IUPAC ambiguity codes [15] for mismatched bases. Sealer records these generated sequences for subsequent insertion into a given scaffold.
To minimize peak memory usage, Sealer performs these local assembly runs serially, such that there is only one Bloom filter loaded at a given time. This implementation is beneficial for processing large genomes, such as P. glauca for which each Bloom filter instance requires 40GB RAM. Further, it allows a subtractive procedure, where we eliminate successfully closed sequence gaps from the input of subsequent iterations, minimizing CPU run time.
-
Step 3: Updating the draft assembly. The scaffold IDs and gap start positions recorded during Step 1 are used to match the new sequences to the corresponding gaps. However, the size of a gap and the length of a filled sequence may disagree, potentially shifting gap coordinates as Sealer closes them. To avoid this issue, Sealer processes gaps from 3′ to 5′, attempting to close the right-most gap first, moving left, making its way gap by gap towards the 5′ end of a scaffold. When a gap and new sequence are matched correctly, Sealer removes the N bases and the flanking nucleotides in the selected window, and replaces them by the newly assembled sequence. The flanking nucleotides are also replaced since the assembly process may have modified portions of these sequences, correcting micro-misassemblies near the end of the flanking sequence by adjusting them so that they are concordant with the underlying read set. This may also occur if there are alternative solutions to the original assembly problem, as in the case of polyploid genomes with allelic variations.
Experimental Data
We used five datasets representing genomes of varying size (from 5 Mbp to 20 Gbp) and complexity (bacterial to conifer) to assess the scalability and performance of Sealer over a range of conditions (Table 1). All draft assemblies were generated using ABySS using optimized assembly parameters. Specifically, we downloaded E. coli K-12 substr. DH10B (~5Mbp) Illumina HiSeq 2000 paired-end reads from the Sequence Read Archive (SRA) (SRA: SRR959238). We used the reference genome [GenBank: NC_010473.1] from NCBI to assess the accuracy of the filled gaps. We closed gaps in this draft assembly with Sealer using seven k-mer lengths (k = 90 to 30 bp, decrementing by 10), and using the Konnector parameters -B 3000 -F 5000 -P 10. We obtained experimental S. cerevisiae S288c (12Mbp) data from the European Nucleotide Archive. The corresponding reference was downloaded from NCBI [GenBank:GCF_000146045.2]. Gaps were closed with Sealer using the same seven k-mer lengths as for E. coli and the Konnector parameters -B 3000 -F 700 -P 10. Experimental C. elegans (~100 Mbp) paired-end reads were obtained from the SRA. The latest version of the reference genome was acquired from WormBase [WormBase:WBcel235]. Sealer closed gaps in the draft assembly using 64 k-mer lengths (k =100 bp, and k-mer lengths between 97 and 35 bp) and the Konnector parameters -B 3000 -b 1200 M -F 700 -P 20. Sequence reads for H. sapiens (3.3 Gbp) individual NA19238 from the 1000 Genomes Project were obtained from the SRA, and the human genome reference GRCh38 obtained from Genome Reference Consortium. Gaps were closed with Sealer using 31 k-mers (250 – 130 bp, decrementing by 10 and 125 – 40 bp, decrementing by 5), and the parameters for Konnector were -B 1000 -F 700 -P10. P. glauca (white spruce) V3 draft assembly (20.8 Gbp) was downloaded from NCBI [GenBank:GCA_000411955.3). We ran Sealer on this dataset using two k-mer lengths (96 and 80 bp) while using the Konnector parameters -B 300 -F 700 -P 10.
Comparison to other gap-closing tools
Sealer was compared to two similar tools: GapFiller (v1.10) [10] and GapCloser (v1.12), the latter in the SOAPdenovo2 package [11]. Default settings were used for both tools in our tests, maximizing the number of compute threads, when needed (−t 16 for GapCloser on the human data set). Smaller datasets (<100 Mbp) were included in the assessment of Sealer to accommodate GapCloser and GapFiller, which have high memory and runtime requirements, respectively.
The two were also tested on the 3-gigabase H. sapiens draft assembly, but GapFiller was manually stopped after running for over 350 h (approximately 14 days) without completion or output. Neither one of the tools were used on P. glauca, based on their compute resource requirements on the high-coverage (71-fold) H. sapiens data.
Machine specifications
All Sealer runs were benchmarked on a 12-core computer running CentOS 5.4 with two Intel Xeon X5650 CPUs @ 2.67 GHz and 48 GB RAM. All GapFiller and GapCloser runs were performed on a machine using CentOS 5.10 with 16 cores @ 2.13GHz, 125 GB RAM with the exception of the GapCloser run on the H. sapiens data. For that we used a machine running CentOS 5.9 with 16 cores @ 2.13 GHz and 236 GB RAM to accommodate the high memory requirement of GapCloser.
Assessment of performance
To ensure consistent reporting of gap statistics, a script was developed that counts the number of regions with N bases in an assembly. This script was used to calculate the number of gaps before and after processing a draft assembly. A gap was defined as having one or a contiguous group of N bases. We used Exonerate (v2.4.0) [16] to analyze the accuracy of each tool. Sequence alignments were used to calculate the average sequence similarities (i.e. percentage of matching bases out of total bases in the query without penalizing ambiguity codes). Inserted sequences of closed gaps, rather than entire assembly scaffolds, were aligned to the corresponding reference (exonerate --percent 95). With Sealer, this was a straightforward process because of the comprehensive output it provides. Along with the new draft assembly and a log file describing specific results of each k run, Sealer also outputs a FASTA-formatted file of all the newly generated sequences (flanking sequences and new gap-filled sequence), which includes the original scaffold ID and gap position. In addition, there is an option to output a fasta file of gap-flanking nucleotide sequence pairs (Sealer parameter --print-flanks). In contrast, neither GapCloser nor GapFiller output a file of newly inserted sequences. Therefore, we had to develop a pipeline to identify and extract these novel bases for assessment and benchmarking the performance between all three tools. The pipeline begins by generating a file of all gap-flanking sequence pairs (100 bp each) found in the original assembly. It performs an Exonerate alignment using these flanking sequence pairs and a gap-filled assembly as query and target, respectively. This returns the coordinates of flanking sequence pairs within the gap-filled assembly. Using this information, the assessment pipeline extracts the bases found between each flanking sequence pair, and aligns them to the reference genome to determine sequence similarity. This pipeline was used on all three tools to maintain consistency in our analyses. We measured the runtime of Sealer using the UNIX command ‘time’. We used the Python script Memusg [https://github.com/jhclark/memusg] to benchmark peak memory usage. In addition to these analysis scripts, gap-closed assemblies of E. coli were manually inspected. Using parts of the sequence assessment pipeline described above, newly generated sequences and their flanking sequences were extracted from gap-closed assemblies. These sequences were aligned to the reference using Exonerate (exonerate --model affine:local [new sequence] [reference] --percent 95 --ryo Percent Identity: %pi Percent Similarity %ps). The alignments of sequences produced by Sealer were compared with the alignments of sequences generated by the other two tools. Indels (insertions and deletions), mismatches and ambiguity codes were noted. We further analyzed the quality of the new draft assemblies using QUAST [17]. Additionally, gene annotations (sources listed in Table 1), allowed us to determine whether the gaps closed by each tool spanned genic regions.
Results and discussion
We benchmarked Sealer, GapFiller [10] and GapCloser [11] on five datasets across a broad spectrum of genome assembly sizes (~5 Mbp to 20 Gbp). The comparators were chosen based on overall performance and resource requirements, as recently evaluated [10, 17]. These studies considered several other tools that we did not include in our comparisons. IMAGE [18] was not considered based on its prohibitively high processing time [10].
Gap-closing capability was similar between GapFiller and Sealer (Fig. 1), with Sealer outperforming the former in two out of three small draft genomes (<1 Gbp). GapCloser never achieved a full gap closing success rate above 50 % in any of the datasets tested. We note that GapFiller and GapCloser both have the ability to resolve some of the unambiguous bases within a gap, even when the gap is not completely closed. In contrast, Sealer produces an all-or-none gap closing output. A summary of the comparison results are presented in Table 2.
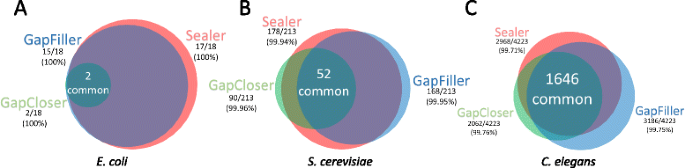
We took advantage of the small size of E. coli to manually inspect every sequence produced by Sealer, GapCloser and GapFiller. We aligned these sequences to the reference genome to determine percent similarities. As shown in Additional file 2: Table S1, Sealer closed 17 of the 18 gaps while GapCloser and GapFiller fully closed 2 and 15 gaps, respectively. Out of the 17 new sequences output by Sealer, 15 had 100 % similarity to the reference genome. GapCloser also had 100 % similarity, but only from its two fully closed sequences. GapFiller performed similarly to Sealer, obtaining 100 % similarity for all but 2 gaps. The gaps commonly closed by all 3 tools (n = 2) comprised the same base sequence. Furthermore, the sequences of commonly closed gaps between Sealer and GapFiller were the same, with the exception of two gaps. Three sequences produced by Sealer contained ambiguity codes, consistent with the ability of Konnector to report alternate bases when multiple assembly paths are possible.
For S. cerevisiae, of the 52 commonly closed gaps (out of 213), percent similarities to the reference genome are 99.94 %, 99.96 % and 99.95 % for Sealer, GapCloser and GapFiller respectively. Sealer successfully closed 178 of the 213 gaps in this dataset, outperforming the other tools.
For C. elegans, 1646 gaps were detected as commonly closed. Sequence similarities were reported as 99.71 %, 99.76 % and 99.75 % for Sealer, GapCloser and GapFiller respectively. These findings (summarized in Table 2 and Fig. 2) show that the accuracy of the gaps closed by the three tools are comparable, while the success rate of Sealer is the highest in all but one experiment.
Identity of closed gaps by Sealer and two leading gap-filling applications. Venn diagrams depict the overlap of gaps closed between each tool for the a) E. coli, b) S. cerevisiae and c) C. elegans datasets. The sizes of individual circles represent the number of gaps closed relative to the other tools. Overlapping closed gaps were approximated using the assessment pipeline described in Section 2.5 and depicted using the online VennDiagram.tk tool [http://www.cmbi.ru.nl/~timhulse/venn/]
We note that the peak memory usage of GapCloser is up to one hundred times higher compared to Sealer (Table 2 and Additional file 3: Figure S2, eg. 101 GB vs 1.35 GB for C. elegans), and memory requirements are similar between Sealer and GapFiller. For smaller genomes, GapCloser had the fastest run times, but consistently closed less gaps compared to the other two tools. GapFiller had the slowest run time for all the experiments. When closing gaps in the H. sapiens draft assembly, GapFiller was left running for over 353 h (~2 weeks) before we manually killed the process. GapCloser was able to complete this dataset in three and a half days using 178.1 GB RAM, while GapFiller was still processing 19 % of the paired-end reads at the time of its termination, with no partial assembly output. Sealer closed 120,676 of the 237,406 human draft assembly gaps, a success rate of 50.8 %, in less than 30 h (~1 day, 5 h) compute time. Compared to GapFiller, Sealer used ~8 times less memory to close marginally more gaps. Likewise, Sealer processed the colossal white spruce 20 Gbp draft assembly in 26.2 h using Bloom filters for two values of k, and achieved a gap-closing success rate of 13.8 %, closing 399,476 of the 2,894,274 gaps in the NCBI V3 draft assembly.
The results reported by QUAST (Additional file 4: Table S2) further supports the ability of Sealer to produce quality draft assemblies. With the exception of the C. elegans assemblies, the resulting Sealer gap-filled assemblies are contiguous, even when factoring mis-assemblies, as the NGA50 length metric suggests. This indicates that the mis-assemblies in the Sealer E. coli and S. cerevisiae assemblies tend to be on shorter scaffolds. In all genomes under study, Sealer was marginally superior in its ability to close gaps located within genes compared to the other tools. Average additional number of complete genes recovered by Sealer compared to other applications for E. coli, S. cerevisiae and C. elegans is 8.5 +/− 6.4, 10.5 +/− 12.0 and 20.0 +/− 1.4 in that order. We speculate that, when measuring number of Ns per 100 kbp, QUAST is counting ambiguity codes as well, since no N bases exist in the contig files submitted to the analysis tool.
Leading gap-filling applications, namely GapFiller and SOAPdenovo’s GapCloser resolve gaps by short read sequence alignments. Current implementations of this paradigm have similar gap-filling efficiencies on small bacterial genomes [10], which is comparable to that of Sealer. They however, have difficulty scaling to larger genomes such as the whole human genome and sizes beyond. This will become increasingly more important as the data throughput from sequencing instruments continues to swell, and researchers undertake more de novo sequencing projects of large genomes. For such projects, even though it is possible to process assembly scaffolds in smaller batches on a compute farm, doing so would result in an overhead that is often impractical and requires specialized hardware and bioinformatics know-how. While our manuscript was in review we became aware of a promising, but not yet scalable, graph-based gap filling algorithm [19], which was reported at the time to close 28 % more gaps in bacterial genome assemblies when compared to GapFiller and GapCloser.
Sealer uses a light-weight Bloom filter de Bruijn graph assembler as its core assembly algorithm. This gives Sealer a few advantages over alternative tools. 1) The entire 1.35 Tbp P. glauca read set [13] easily fits on a 48 GB RAM computer, a system that is now accessible to most labs. 2) The de novo assembly method Sealer uses has the advantage of generating multiple paths through a k-mer graph, which could be used to effectively capture allelic differences and sequence variants, both key features to genetics and cancer studies. In contrast, GapCloser and GapFiller use coverage information and a threshold score to determine which consensus base is used in the case of a discrepancy, effectively stripping that information. 3) The fast bi-directional de Bruijn graph assembly strategy, specifically allows one to exhaustively assemble k-mers across gaps using a comprehensive range of k values. We applied this strategy to the C. elegans dataset, testing 64 different k values (thus iteratively building 64 Bloom filters). Although this impacted its run time, Sealer was still four times faster than GapFiller while closing above 70 % of the gaps, a yield comparable to that obtained by the latter. Further, because the Sealer run time is low and its memory footprint is relatively small, one could envision building Bloom filters with additional data from the same sample, or even third-party data from same-species, to produce a mosaic assembly in a manner similar to the human genome reference.
When running individual Sealer runs at unique values of k on 250 bp human experimental WGS read data, we find that k = 80 is more successful at closing gaps. When considering the entire k spectrum, being more permissive on the maximum number of assembly paths (from –P 2 to –P 10) increases gap closure by 9.8 % overall (Additional file 5: Figure S3A). Generally speaking, k-mers varying in size from 60 to 220 bp were all suited to close gaps in the human draft assembly, and gaps of equivalent sizes tend to close in a k-independent manner, with a slight constriction of gap length distribution with decreasing k (Additional file 5: Figure S3B). This is not surprising since Konnector achieves maximum efficiency on fragments < 1 kbp [12]. On a practical note we recommend, whenever possible, exploring a wide range of k, typically from the read length L to k = 40, which is the practical lower limit of k for Konnector. For human, we ran Sealer iteratively by exploring 31 k-mers from 250 to 40 nucleotides, and still completed the run in less than half the time compared to the next best application (Table 2). When working with de Bruijn graphs, shorter k-mers yield more tangled graphs but are useful when read coverage is inadequately low. On the flip side, the use of longer k-mers help disambiguate repeats and tangles in the graph when the coverage is sufficient, and are expected to help resolve gaps that arise due to repeats [20]. We find that, when used iteratively, both large and short k-mers close a similar number of gaps in the human data set (Additional file 6: Table S3). A combination of k-mers used in iterative cascading runs is thus warranted since, clearly, lower k values close more than half of the gaps that were not successful at larger values of k. Also, larger k-mer values tend to close larger gaps and overlaps. Generally, the first k-mer utilized closes the most gaps in iterative runs (Additional file 7: Figure S4A), which is why running Sealer first with large k values that can more readily resolve repeats is recommended. We profiled the repeat content in gaps [21] that were uniquely closed at a specific k-value during the iterative Sealer run on the human assembly. We observe that gaps with SINEs are preferentially closed at a shorter k while those comprised of simple repeats, LINEs and Satellites are preferentially closed with larger k-mers, which is intuitive. Overall, we do not find many gaps harboring low complexity regions, small RNA, and unclassified repeats, suggesting that we may have limited success in closing gaps comprising these features (Additional file 7: Figure S4B). We have randomly sampled 350 gaps that we could not close after the Sealer iterative k runs on the human assembly draft. We observe that 95.6 % +/− 2.0 % of those are due to repeats in the hg19 reference human genome, as indicated by the corresponding regions harboring lower case bases in the reference genome. The +/− 2.0 % interval refers to the 95 % confidence interval of this estimated rate.
GapCloser and GapFiller will resolve base ambiguities as they iterate through a subset of sequences even when they cannot fully close a gap. Sealer on the other hand provides an all-or-nothing output, not reporting partially filled gaps when the number of possible reconstructions between flanking gap sequences exceed a user-defined threshold or when assembly paths are obscured. As a result, GapCloser and GapFiller results may have fewer Ns per 100 kbp (Additional file 4: Table S2). However, we note that remaining gaps still require design and execution of finishing experiments, and arguably fewer gaps would be desirable over more but slightly shorter gaps. Also, because GapCloser and GapFiller do not report their partially closed gaps, we were not able to test their accuracy.
Conclusions
Finishing genomes has been relevant since sequencing H. influenzae, the first shotgun genome assembly [22] and, more than ten years after publishing of the first human genome draft, we still do not have a complete assembly [23]. There have been few reasons for that, including difficulty in cloning, sequencing and assembling heterochromatin, as well as the lengthy, tedious nature of finishing work. But the prohibitive analysis cost is one of the main reasons why we do not completely finish genomes today. With DNA sequencing affordability on the rise [4], shotgun assembly of large genomes (>1 Gbp) is increasingly becoming routine in research laboratories, and widespread uptake in the clinic is anticipated. Consequently, obtaining better draft genomes is a common goal of all de novo sequencing projects. Further, de novo assembly and sequence finishing is also finding applications in many re-sequencing projects, especially in oncogenomics projects where comprehensive sequencing and nucleotide-level resolution informs clinical intervention [24]. Sealer is a scalable gap-filling software expected to be an indispensable addition to the genome finishing toolkit, and with broad application to on-going finishing efforts.
Whereas obtaining 100 % completion is unlikely without at least some computer-assisted manual finishing and labour-intensive PCR work, Sealer brings human genome finishing and finishing of colossal genomes such as that of the 20 Gbp white spruce one-step closer.
Availability and requirements
Project name: Sealer
Project home page: https://github.com/bcgsc/abyss/tree/sealer-release
Operating system(s): UNIX
Programming language: C++
Other requirements: Boost C++ library (headers only) and Google sparsehash library
License: GNU GPL
Any restrictions to use by non-academics: license needed
Abbreviations
- ABySS:
-
Assembly By Short Sequences
- WGS:
-
Whole Genome Shotgun
- IUPAC:
-
International Union of Pure and Applied Chemistry
- RAM:
-
Random Access Memory
- Gbp:
-
Giga base pairs
- CPU:
-
Central Processing Unit
References
Simpson JT, Wong K, Jackman SD, Schein JE, Jones SJ, Birol I. ABySS: a parallel assembler for short read sequence data. Genome Res. 2009;19:1117–23.
Stein LD. The case for cloud computing in genome informatics. Genome Biol. 2010;11:207.
1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, et al. A map of human genome variation from population-scale sequencing. Nature. 2010;467:1061–73.
Mardis ER. The $1000 genome, the $100,000 analysis? Genome Med. 2010;2:84.
Cancer Genome Atlas Research Network. Genomic and Epigenomic Landscapes of Adult De Novo Acute Myeloid Leukemia. N Engl J Med. 2013;368:2059–74.
Pugh TJ, Morozova O, Attiyeh EF, Asgharzadeh S, Wei JS, Auclair D, et al. The genetic landscape of high-risk neuroblastoma. Nat Genet. 2013;45:279–84.
Roberts KG, Morin RD, Zhang J, Hirst M, Zhao Y, Su X, et al. Genetic Alterations Activating Kinase and Cytokine Receptor Signaling in High-Risk Acute Lymphoblastic Leukemia. Cancer Cell. 2012;22:153–66.
Yip S, Butterfield YS, Morozova O, Chittaranjan S, Blough MD, An J, et al. Concurrent CIC mutations, IDH mutations, and 1p/19q loss distinguish oligodendrogliomas from other cancers. J Pathol. 2012;226:7–16.
Hunt M, Newbold C, Berriman M, Otto TD. A comprehensive evaluation of assembly scaffolding tools. Genome Biol. 2014;15:R42.
Boetzer M, Pirovano W. Toward almost closed genomes with GapFiller. Genome Biol. 2012;13:R56.
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler.Gigascience. 2012;1:18.
Vandervalk BP, Jackman SD, Raymond A, Mohamadi H, Yang C, Attali DA, et al. Konnector: Connecting paired-end reads using a bloom filter de Bruijn graph. Bioinformatics Biomedicine (BIBM). 2014. doi:10.1109/BIBM.2014.6999126.
Birol I, Raymond A, Jackman SD, Pleasance S, Coope R, Taylor GA, et al. Assembling the 20 Gb white spruce (Picea glauca) genome from whole-genome shotgun sequencing data. Bioinformatics. 2013;29:1492–7.
Chikhi R, Rizk G. Space-efficient and exact de Bruijn graph representation based on a Bloom filter. Algorithms Mol Biol. 2013;8:22.
Cornishbowden A. Nomenclature For Incompletely Specified Bases In Nucleic-Acid Sequences - Recommendations 1984. Nucleic Acids Res. 1985;13:3021–30.
Slater GS, Birney E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics. 2005;6:31.
Gurevich A, Saveliev V, Vyahhi N, Tesler G. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29:1072–5.
Tsai IJ, Otto TD, Berriman M. Improving draft assemblies by iterative mapping and assembly of short reads to eliminate gaps. Genome Biol. 2010;11:R41.
Salmela L, Sahlin K, Mäkinen V, Tomescu AI. Gap Filling as Exact Path Length Problem. In: Przytycka TM, editor. Research in Computational Molecular Biology. Lecture Notes in Computer Science Volume 9029. Warsaw: Springer International Publishing; 2015. p. 281–292.
Birol I, Jackman SD, Nielsen CB, Qian JQ, Varhol R, Stazyk G, et al. De novo transcriptome assembly with ABySS. Bioinformatics. 2009;25:2872–7.
Smit, AFA, Hubley, R & Green, P. RepeatMasker Open-4.0. 2013–2015 http://www.repeatmasker.org.
Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science. 1995;269:496–512.
Genovese G, Handsaker RE, Li H, Kenny EE, McCarroll SA. Mapping the human reference genome’s missing sequence by three-way admixture in Latino genomes. Am J Hum Genet. 2013;93:411–21.
Jamshidi F, Pleasance E, Li Y, Shen Y, Kasaian K, Corbett R, et al. Diagnostic value of next-generation sequencing in an unusual sphenoid tumor. Oncologist. 2014;19:623–30.
Acknowledgements
The authors thank the funding organizations, Genome Canada, British Columbia Cancer Foundation, and Genome British Columbia for their partial support. This research report was also partly supported by the National Institutes of Health under Award Number R01HG007182. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or other funding organizations.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
R.L.W. and I.B. designed the research; R.L.W. and D.P. prototyped the application; D.P., B.P.V., T.R. and S.J. developed the software. D.P. and R.L.W. analyzed the data and made the figures; D.P., R.L.W. and I.B. wrote the manuscript. All authors discussed the results and commented on the manuscript. All authors read and approved the final manuscript.
Daniel Paulino and René L. Warren contributed equally to this work.
Additional files
Additional file 1: Figure S1.
Sealer gap-closing pipeline. PDF file depicting the steps to gap-filling genome draft assemblies with Sealer.
Additional file 2: Table S1.
Table of gaps closed in the E. coli K-12 genome. PDF file presenting an in-depth gap sequence analysis in E. coli.
Additional file 3: Figure S2.
Compute resource required. PDF file showing benchmarking results of Sealer, SOAPdenovo GapCloser and GapFiller for closing gaps in draft genome assemblies.
Additional file 4: Table S2.
Table detailing a quality assessment of gap-filled assemblies. PDF file reporting a QUAST analysis of gap-closed E. coli, S. cerevisiae and C. elegans draft genome assemblies.
Additional file 5: Figure S3
Individual Sealer H. sapiens runs at unique values of -k and –P. (PDF 126 kb)
Additional file 6: Table S3
Closing gaps in a human genome draft assembly using Sealer with different k value ranges. (PDF 54 kb)
Additional file 7: Figure S4.
Closing gaps iteratively in a cascading fashion at various values of k in the E. coli, S. cerevisiae, C. elegans, H. sapiens and P. glauca draft genomes.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver (https://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Paulino, D., Warren, R.L., Vandervalk, B.P. et al. Sealer: a scalable gap-closing application for finishing draft genomes. BMC Bioinformatics 16, 230 (2015). https://doi.org/10.1186/s12859-015-0663-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12859-015-0663-4