Figure 4
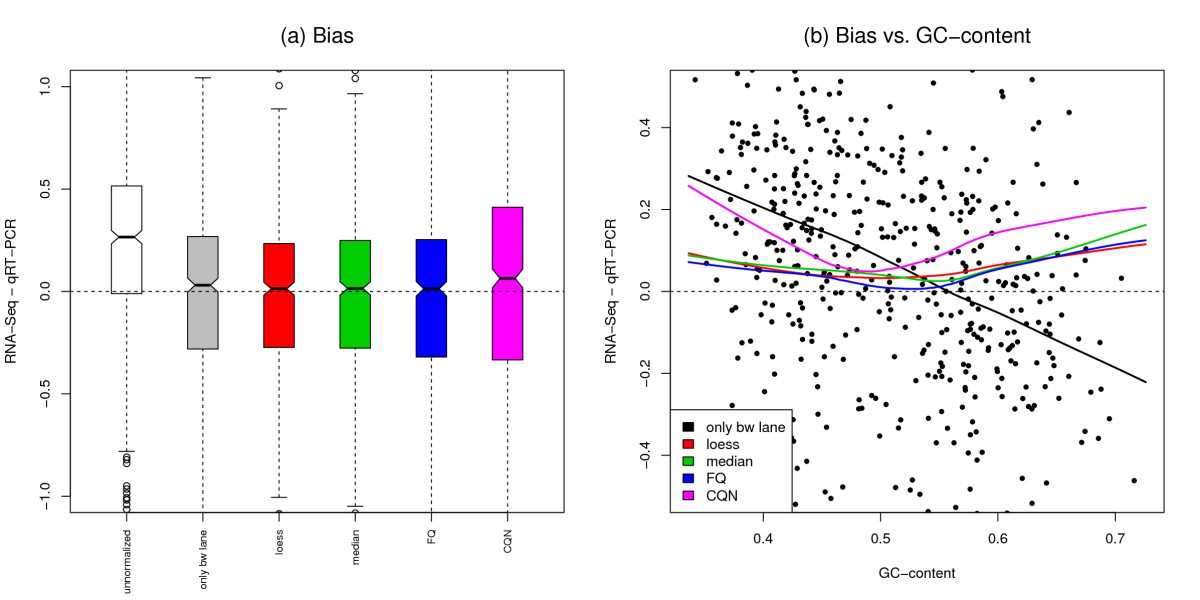
MAQC-2 dataset: Bias in fold-change estimation. Bias in UHR/Brain expression log-fold-change estimation for different RNA-Seq normalization procedures, where bias is defined as the difference between the estimates from RNA-Seq and qRT-PCR for 638 genes assayed by both technologies. Panel (a): Boxplots of bias in log-fold-change estimates. Our three proposed normalization procedures reduce bias, while CQN tends to overestimate the UHR/Brain fold-change. Panel (b): Dependence of bias on GC-content. The points correspond to bias after only FQ between-lane normalization, the curves are lowess fits of bias vs. GC-content for different normalization procedures. There is still substantial dependence of bias on GC-content after CQN.