Figure 11
From: MUSCLE: a multiple sequence alignment method with reduced time and space complexity
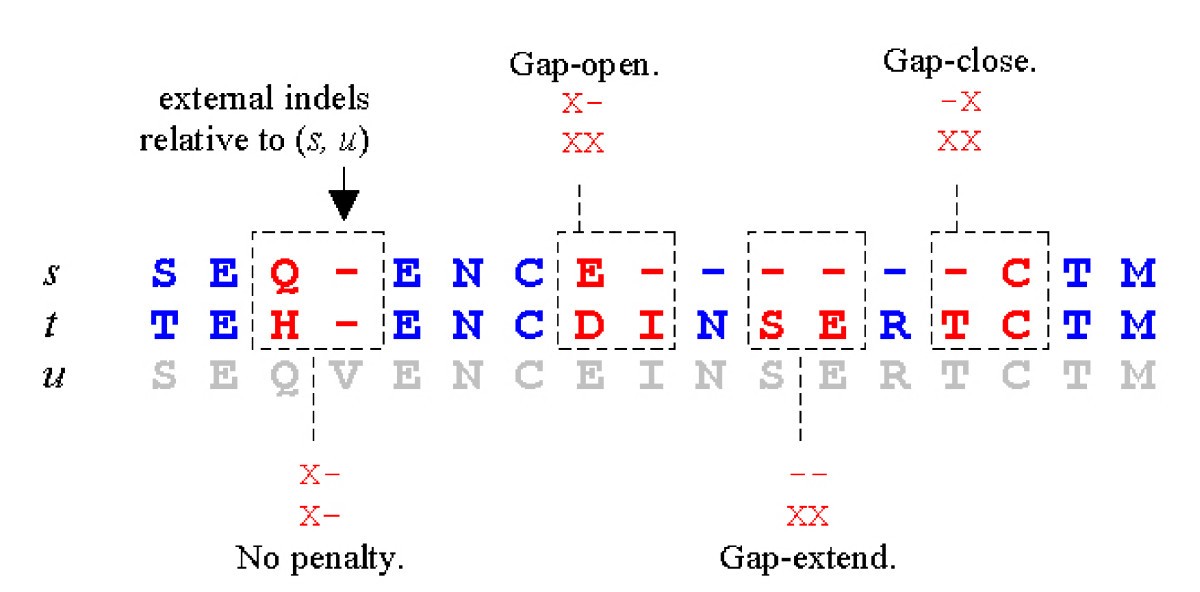
Dimers in the {X,-} alphabet. Gap penalties for the sequence pair (s, u) can be computed be considering all aligned pairs of dimers in the alphabet {X,-}, where X is any amino acid and - is the usual indel symbol. Four cases are highlighted. Note that an aligned pair of identical dimers never contribute a gap penalty as any indels in the dimers are necessarily external, as in the left-most example.