Figure 8
From: Fast index based algorithms and software for matching position specific scoring matrices
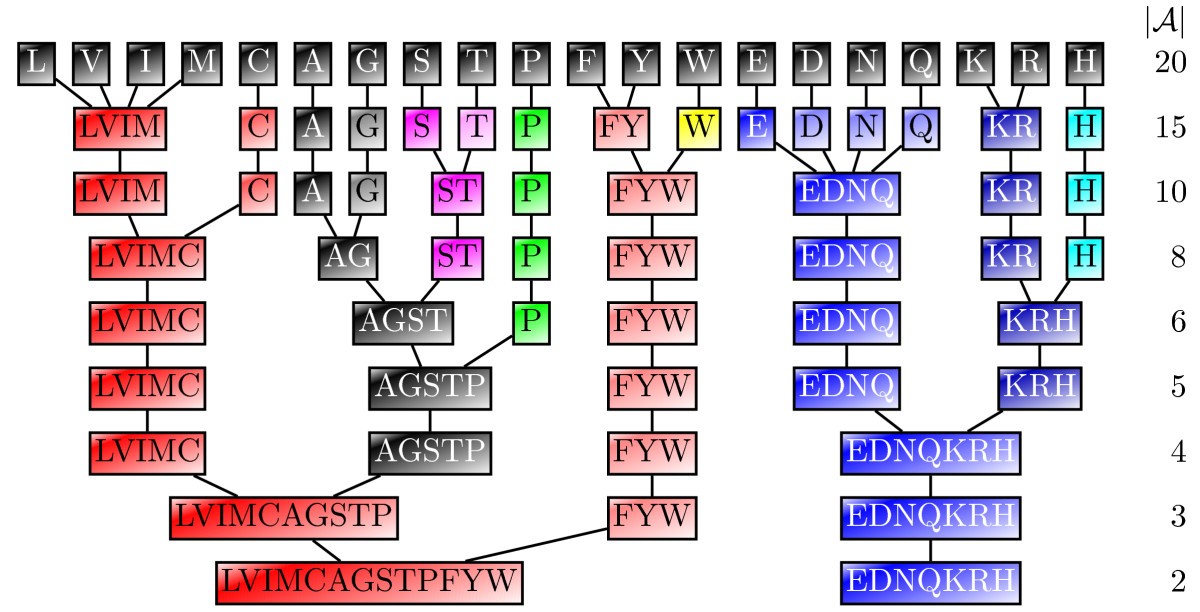
Schemes for amino acid alphabet reduction. Reduction of the amino acid alphabet into smaller groups. Amino acid pairs are iteratively grouped together based on ther correlations ca,b(see text for the definition of ca,b), starting with the most correlated pairs, until al amino acids are divided into the desired number of groups. Here we used BLOSUM50 similarities for the determination of ca,b. Observe that, hydrophobic amino acids, especially (LVIM) and (FYW) are conserved in many reduced alphabets. The same is true for the polar (ST), (EDNQ), and (KR) groups. The smallest alphabet contains two groups that can be categorized broadly as hydrophobic/small (LVIMCAGSTPFYW) and hydrophilic (EDNQKRH).