- Database
- Open access
- Published:
Propedia: a database for protein–peptide identification based on a hybrid clustering algorithm
BMC Bioinformatics volume 22, Article number: 1 (2021)
Abstract
Background
Protein–peptide interactions play a fundamental role in a wide variety of biological processes, such as cell signaling, regulatory networks, immune responses, and enzyme inhibition. Peptides are characterized by low toxicity and small interface areas; therefore, they are good targets for therapeutic strategies, rational drug planning and protein inhibition. Approximately 10% of the ethical pharmaceutical market is protein/peptide-based. Furthermore, it is estimated that 40% of protein interactions are mediated by peptides. Despite the fast increase in the volume of biological data, particularly on sequences and structures, there remains a lack of broad and comprehensive protein–peptide databases and tools that allow the retrieval, characterization and understanding of protein–peptide recognition and consequently support peptide design.
Results
We introduce Propedia, a comprehensive and up-to-date database with a web interface that permits clustering, searching and visualizing of protein–peptide complexes according to varied criteria. Propedia comprises over 19,000 high-resolution structures from the Protein Data Bank including structural and sequence information from protein–peptide complexes. The main advantage of Propedia over other peptide databases is that it allows a more comprehensive analysis of similarity and redundancy. It was constructed based on a hybrid clustering algorithm that compares and groups peptides by sequences, interface structures and binding sites. Propedia is available through a graphical, user-friendly and functional interface where users can retrieve, and analyze complexes and download each search data set. We performed case studies and verified that the utility of Propedia scores to rank promissing interacting peptides. In a study involving predicting peptides to inhibit SARS-CoV-2 main protease, we showed that Propedia scores related to similarity between different peptide complexes with SARS-CoV-2 main protease are in agreement with molecular dynamics free energy calculation.
Conclusions
Propedia is a database and tool to support structure-based rational design of peptides for special purposes. Protein–peptide interactions can be useful to predict, classifying and scoring complexes or for designing new molecules as well. Propedia is up-to-date as a ready-to-use webserver with a friendly and resourceful interface and is available at: https://bioinfo.dcc.ufmg.br/propedia
Background
Peptides are short chains of amino acid residues connected by peptide bonds that act in cell signaling and as immune modulators, among other important functions. It is estimated that between 15 and 40% of all protein–protein interactions in cells are mediated by these molecules [1]. Additionally, peptides are structurally diverse, versatile, induce low resistance with limited nontarget activity and can be modulated to interact with specific cellular targets, making them good therapeutic agents [2]. However, their short half-life and poor oral bioavailability has discouraged the search for peptides as therapeutics in the past [3].
With the recent emergence of new synthetic approaches that permit changes in the biophysical and biochemical properties of peptides, these molecules are once again being considered as drug candidates [4,5,6]. In fact, over 60 peptide drugs have been approved in major pharmaceutical markets and hundreds of others are in active clinical development at the moment [3]. Peptide-like inhibitors are used as well to treat cancer, diabetes, and autoimmune diseases and have high success rates in commercial development [7]. Multiple next-generation drug candidates (derived from exenatide, a synthetic form of a natural 39-amino acid peptide secreted by Heloderma suspectum), have been proposed as therapeutic agents for type 2 diabetes mellitus [3].
Understanding the structure and recognition of protein–peptide complexes may aid the design of novel peptides and peptide-based compounds for drug development or biotechnological purposes. Databases of protein–peptide complexes can pave the way for the analysis and comprehension of the mechanisms of protein–peptide recognition. There are several peptide databases, with varied purposes, as databases of bioactive peptides [8], antimicrobials [9], cell penetrating peptides [10], hemolytic peptides [11], etc. [8]. Here, we briefly review some representative examples of protein–peptide databases.
London and colleagues [12] in 2010 proposed PeptiDB, comprising 103 high-resolution peptide-protein complex structures. It was proposed as a nonredundant set of high resolution complexes to investigate the structural bases of interactions between proteins and peptides and to improve understanding binding strategies for short peptides (5–15 residues).
Also in 2010, Vanhee et al. [13] devised PepX, comprising protein–peptide complexes clustered based on binding interfaces. It was updated in 2014 for the last time (505 unique protein–peptide interface clusters from 1431 complexes) and is not available anymore.
Das et al., in turn, proposed PepBind [14] in 2013 as a curated set of 3100 protein–peptide complexes clustered according to structure determination methods and manually curated for cellular activity of complexes. The authors mentioned that there was a web interface but it seems to no longer be available.
More recently, in 2018, Frappier et al. [15] presented PixelDB a database that comprises 1966 non-redundant high-resolution complexes. Entries are clustered based on structural similarities of receptors and then on binding modes. The authors claim to identify conserved peptide core structural motifs. We found a version of this database on GitHub updated 3 years ago.
Wen et al. [16] released PepBDB also in 2018 and this database is available through a web interface and for download. It contains 13,299 complexes and was last updated in March 2020. The web interface presents the whole list and an individual interactive visualization of the 3D interface and a 2D plot of hydrogen bonds and hydrophobic interactions using LigPlot [17]. Protein–peptide complexes can be filtered considering sequence features, structure resolution and experimental method.
At the end of 2019, Xu et al. [18] proposed PepPro, a nonredundant benchmarking tool for testing peptide-protein docking algorithms composed of only 89 complexes. For 58 complexes, the unbound protein structures are available, which is useful for evaluating to what extent docking algorithms can accommodate binding-related protein conformational changes.
In summary, a variety of databases have been proposed to explore and increase the understanding of protein–peptide interactions. Nevertheless, despite their relevant contributions when released, most of them are obsolete and/or no longer supported. Among those mentioned, PepBDB is the most comprehensive, as it contains approximately 13,000 complexes. In addition, it is the only one that provides features for binding mode analysis.
To fill these gaps, aiming at automatically collecting a broad and up-to-date data-set of protein–peptide complex structures as a useful resource for diverse peptide studies, we propose Propedia. This database is a comprehensive, general purpose and up-to-date protein–peptide resource that contains over 19,000 high-resolution structures from the Protein Data Bank (PDB) segmented in clusters to reduce redundancy if desired. Structures of complexes have been organized, facilitating search and visualization by different criteria such as PDB id, sequence similarity, peptide classification, source organism, binding area, molecular weight, aromaticity, instability index, isoelectric point, and hydrophobicity, among other computed data. These clusters not only help accommodate redundancy in the database but also allow comparisons among sequences, interfaces, interactions and functions. Therefore, Propedia is a comprehensive and powerful tool for structural studies of protein–peptide recognition, support for construction of training and test data sets for docking and scoring approaches, and facilitation of peptide rational design.
Propedia was inspired by our previous work on defense of plants against insects and pathogens. Soybean, when injured by the caterpillar Anticarsia gemmatalis Hübner, produces the Kunitz trypsin inhibitor (KTI) and the Bowman-Birk inhibitor (BBI), which impede protease-catalyzed degradation in the insect gut [19, 20]. Based on these inhibitors that are naturally produced by soybean, we are interested in proposing peptide or mimetic peptide molecules to inhibit the proteases of the caterpillar gut. We believe these molecules have the potential to be used in the ecological control of this pest insect. We formerly designed peptides manually, with the support of certain bioinformatic tools. Now we are investing in the development of automatic tools to support this process, such as ppiGReMLIN [21]. In this context, Propedia aims to deliver a comprehensive data set of experimental protein–peptide complexes organized in three types of clusters based on : (1) sequence similarity; (2) interface structure; and (3) protein–peptide binding site. It permits analysis of structures under different perspectives, supporting the detection of potential peptides for interacting with a target of interest, for example, peptides that are likely to inhibit proteases of the caterpillar gut. It is important to note that our database is not specific to soybean and its insect pest Anticarsia gemmatalis Hübner and can be applied to other data sets involving protein–peptide complexes.
Construction and content
In this section, we detail the project decisions and the design process followed to build Propedia as well as the contents of the database.
Database construction
We used the following criteria to retrieve PDB entries: (1) structures composed by two or more chains, (2) one chain with at least 2 and no more than 50 residues (for peptides), and (3) structures solved by NMR or X-ray crystallography with resolution below 2.5 Å. The present release is composed of 19,813 complexes (May 02, 2020). We developed in-house Python scripts and the Biopython library [22] to extract PDB data and populate the database. Each file was filtered to remove hydrogen atoms, water molecules, alternative positions [23] and crystallographic artifacts [24].
We identified protein–peptide complexes from the remaining files. Chains with lengths of 2–50 residues were classified as “peptides”. The reason for this choice is to keep Propedia comprehensive comprising the ranges used by the existing databases. Chains with more than 60 residues were classified as “receptors”. This decision was empirical since, by allowing smaller receptors, we observed complexes involving two peptides (or small unstructured proteins).
The protein–peptide interfaces were computed as follows: if there was at least one peptide atom at a distance of 6Å from any receptor atom and the protein–peptide complex had an interface area (greater than 0), then the protein–peptide complex was included in the database. We used the method of Lee and Richards [25] to compute the interface area (IA) and the accessible surface area (ASA). This algorithm returns the surface area of a protein in Å\(^{2}\) and was computed by NACCESS [26] software. The software receives a PDB file as input and returns the ASA of each atom. The IA was calculated using the following equation:
where ASA(A) and ASA(B) are the ASA of the protein (A) and peptide (B), respectively, while ASA(AB) is the protein–peptide complex (AB) ASA. Then, IA is assumed to be the set of atoms that gained solvent accessibility.
With this procedure, we identified 19,813 complexes, including 19,177 from X-ray structures and 636 by NMR. There were peptides missing residues or containing nonstandard amino acid residues or binding with multiple chains describes in Table 1. Peptides bound with multiple receptors may affect both it is structural conformation and those interface residues. Therefore, we removed these complexes, obtaining 5971 protein–peptide complexes and, from now on, we refer to them as the Clusterable Complex Dataset (CCD).
Propedia database schema, presenting the tables, fields and relationships. The complex table (white) is the core of the database and interconnects all the data; pdb entities (blue) including group, pdb_groups and pdb tables; peptide/receptor and organism tables (yellow); cluster tables (green); and alignment tables (orange)
Data collected in previous steps and computed clusters were stored in a MySQL database. The entity-relationship model is depicted in Fig. 1. We have the following entities: pdb, complex, peptide, receptor, organism, cluster (three types: sequence, interface, binding site) and alignment (clustal (peptide sequence), mustang, probis). The group table contains keywords derived from the pdb classification. For example, the Coronavirus Main Proteinase (3CLpro) (PDB id: 1p9u) is classified as ‘Hydrolase/Hydrolase Inhibitor’ and was labeled so as to be included in the groups: ‘Hydrolase’ and ‘Inhibitor’. Alignment tables store data from the results of molecular pair alignment, according to the type of clustering, and therefore have double foreign keys (id_complex1, id_complex2) corresponding with the complex table due to efficiency requirements.
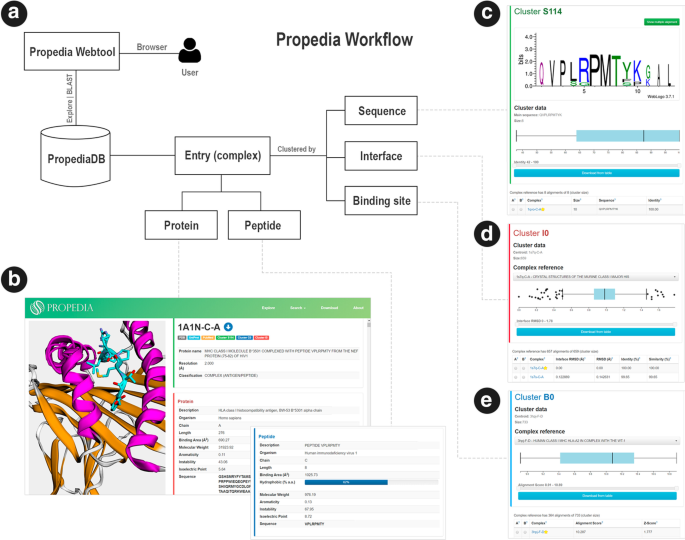
a Propedia scheme. The user accesses Propedia through a browser. Propedia presents each protein–peptide as a complex. Each complex can be associated with a cluster based on sequence, interface or binding site. b Propedia interface. Three-dimensional structure visualization of a complex. Protein is shown as a cartoon (alpha-helix in magenta and beta-strands in orange). The peptide is shown as a cartoon with cyan sticks. Complex information includes receptor features, peptide features, clustering classification and similar complexes. c–e Sequence, interface and binding site, cluster pages. Sequence cluster containing the sequence WebLogo (consensus) and main sequence. Each cluster page has a distribution chart (boxplot), used to filter complexes, according to the attributes used for clustering: sequence identity, iRMSD and alignment score
Clustering
Sequences
We classified peptide sequences using the tool Hammock (1.2.0) [27]. It uses hidden Markov model profiles for peptide sequence clustering and three external tools for multiple alignments, similarity search, and HMM-HMM comparison: Clustal Omega [28, 29], HMMER 3.0 [30], and HHSuite [31]. We ran Hammock using mode ‘full’ with default parameters with the exception of ‘–min_conserved_positions’, which was set to 3, and ‘–count_threshold’, which was set to 300. These values were set empirically. Sequence labels were assigned using Python in-house scripts. CCD was used as input and after the filtering step, Hammock returned 3,495 unique sequences and classified them into 771 clusters and 1074 unique clusters (singletons), totaling 1845 peptide sequence clusters. For each cluster (non singletons) a consensus sequence was generated using the WebLogo tool [32], and the sequence alignment was determined using Clustal Omega [28, 29] to store the sequence identity among the peptides of each cluster. Centroids were identified as the peptides having the same sequence as the main sequence of each cluster.
Interface
Protein–peptide interfaces were aligned with MUSTANG [33], a multiple protein structural alignment tool that superposes structures using the distances of the C-\(\alpha\) coordinates of residues. A pairwise structural alignment was performed using only the protein structures of the CCD. To avoid unfavorable pairwise alignments, we considered only pairs of receptors sharing over 50% sequence identity. A total of 353,545 alignments were performed in parallel in a multicore processor, and the interface RMSD (iRMSD) was calculated from the results. The protein–peptide interface was considered to be all residues within 6 Å of a peptide [34, 35]. In-house Python scripts were developed to create an undirected graph network using the NetworkX (version 1.11) Python library [36]. Nodes representing receptors and the edges (with the iRMSD between them) were added if 75% of the residues that composed the interfaces were aligned and had C\(\alpha\) distance less than or equal to 2 Å. This threshold is the same for PepX [13]. Each connected subgraph from the undirected graph was considered a cluster. Altogether, 535 clusters were formed, plus 1356 singletons, for a total of 1891 non redundant protein–peptide interfaces. Each centroid was defined as the receptor node with the highest degree, and in the case of a tie, the one with the lowest sum of iRMSDs.
Binding sites
We used the ProBiS algorithm [37] to identify similar protein–peptide binding sites. ProBis is a local alignment algorithm that aligns similar binding sites in proteins with dissimilar folds through 3D patterns of physicochemical properties of their surfaces, considering geometrical and functional groups. Functional groups are specific groups of atoms in residues with particular physicochemical properties, which include hydrogen bond acceptors, hydrogen bond donors, acceptor/donors, aromatics and aliphatic groups [38]. ProBiS returns an alignment score for each pairwise alignment. The higher the alignment score, the more similar the binding sites are. It also computes a Z-score, a statistical measure, based on alignment scores of the population. This parameter is calculated using the Karlin-Altschul equation [39]. The input we supplied to Propedia was the CCD and we extracted surface structural patches of each receptor at a distance of 6 Å from the corresponding peptide. Then, we performed a pairwise alignment using ProBiS.
The population mean (\(\mu\)) and population standard deviation (\(\sigma\)) were computed from pairwise alignment scores in the CCD, where \(\mu\) and \(\sigma\) are 1.488 and 4.951, respectively.
We used a similar method to define the clusters based on interfaces. An edge with alignment score, as weight, between two nodes (receptors) was created if the Z-score between them was greater than 1.5. This value was estimated as the point at which the number of clusters starts to increase exponentially. Connected subgraphs defined each cluster, and centroids were selected in the same way we described for previous clusters. Finally, 521 clusters and 945 singletons were generated, totaling 1466 distinct binding sites.
Propedia webserver
The propedia database can be accessed through an interactive webserver implemented in the CodeIgniter PHP framework. Graph visualizations were implemented with the D3.js library (https://d3js.org). Protein–peptide three-dimensional structure visualizations were generated using the 3Dmol.js library [40]. The receptor/peptide sequence search mechanism is based on the blastp tool from NCBI-BLAST+ suite [41, 42] and for the binding site search we use the ProBiS algorithm [37].
Utility and discussion
The Propedia interface (https://bioinfo.dcc.ufmg.br/propedia) is user-friendly, visual and interactive. It allows database searches with several options (Fig. 2a). Each entry in Propedia represents a protein–peptide complex. The web tool allows access to entries through PDB id, which can be followed (optionally) by protein chain id and peptide chain id.
a Structural alignment between 2JF9 and 4IV2. The protein residues were conserved, but the peptide residues were not. b Estrogen receptor alpha LBD in complex with a tamoxifen-specific peptide antagonist (PDB id: 2jf9; peptide chain: Q; protein chain: B). c Estrogen receptor alpha ligand-binding domain in complex with dynamic way-derivative (PDB id: 4IV2; peptide chain: C; protein chain: A)
Propedia’s interface allows searching by pdb, complex id, organism, group (classification keyword), peptide and protein sizes, resolution, protein and peptide sequences (using BLAST), protein binding site (using ProBiS), and similar complexes using different clustering methods. When the user selects a particular complex to analyze, the web page presents the pdb, complex id, resolution, protein/peptide description and organism, and their data, includes chain, length, binding area (Å\(^{2}\)), molecular weight, hydrophobic percent (peptide only), aromaticity, instability index, isoelectric point, and sequence. We enriched Propedia with other relevant information from multiple databases such as UniProt and PubMed: protein chain, length, binding area and sequence information (Fig. 2b).
The Propedia database was built based on a hybrid clustering approach that segments the set of complexes by the following: (1) sequence similarity; (2) interface structure; and (3) protein–peptide binding site. Due to this organization in clusters, users find similar protein–peptides complexes not only by traditional sequence and/or structure conservation but by interactions as well. Interactions, in fact, are essential for molecular recognition. A user can choose among these three different approaches to eliminate redundancy of the data set, if needed.
Comparison with other peptide databases
There are several peptide databases available. Table 2 compares some of their features. Each existing database contributes mainly to a specific piece of biological information. PepX [13] is a protein–peptide interaction database clustered by binding interfaces. It has 1,431 complexes with peptide sizes between 5 and 35 amino acids. PepBind [14] compiles structures, sequences and experimental information for protein–peptide complexes with peptides up to 35 amino acids. PeptiDB [12] comprises only 103 high-resolution complexes with peptides ranging in size from 5 to 15 amino acids. Some of these databases are not being updated and, for others, data are not even available.
PepPro [18], PixelDB [15] and PepBDB [16], on the other hand, are more recent and up-to-date efforts. They aggregate structural data from peptides up to 50 amino acids. PepPro [18] is a benchmark database built specifically for evaluation of protein–peptide docking algorithms. It contains 89 nonredundant complex structures retrieved from 1,198 high-resolution PDB entries with peptide size ranging from 5 to 30 residues. PixelDB [15] contains 1,966 nonredundant protein–peptide structures organized into clusters to provide structural conservation data for peptide binding modes. Finally, PepBDB [16] comprises 12,241 protein–peptide complex structures and their interaction information and is useful for analyzing and benchmarking docking algorithms and scoring functions.
Propedia is a more recent and fully automated database and webserver that will be updated quarterly. It is broader (comprises the entire PDB data-set) and general purpose protein–peptide analysis tool that is ready to collect, filter, clean, and compute several features and cluster data automatically always providing a comprehensive and up-to-date resource. For instance, researchers can already retrieve SARS-CoV-2 proteins along with peptides in Propedia.
Case studies
We designed three case studies using Propedia’s varied features to exemplify possible use cases of the tool and the adjoining database.
Estrogen receptors in complexes with different peptides (2JF9 and 4IV2)
We performed a case study with the estrogen receptor alpha LBD in complex with a tamoxifen-specific peptide antagonist (PDB id: 2jf9; peptide: chain Q; protein: chain B). This is a Homo sapiens protein classified in the PDB in the transcription category. The main objective of this case study was to test if Propedia would be able to find structures with similar binding sites but with different peptide sequences.
We compared the estrogen complex with the crystal structure of the estrogen receptor alpha ligand-binding domain in complex with dynamic way-derivative (PDB id: 4IV2; peptide: chain C; protein: chain A). Although these complexes were classified in the same cluster (B1) considering their interactions, the clusters for sequence was different (Table 3).
a PDB ID: 1lvb; peptide: chain D; protein: chain B; Rosetta score: − 538.306; Distance: 3.5 b PDB ID: 5om5; peptide: chain B; protein: chain A; Rosetta score: − 538.985; Distance: 3.7 c PDB ID: 1lvm; peptide: chain C; protein: chain A; Rosetta score: − 528.398; Distance: 5.5 d the whole set of evaluated peptides
Thus, we aligned the PDB files using the PyMol tool [43] and compared the results manually (Fig. 3). We observed that although the peptide primary structures were different, the peptide \(\alpha\)-helix folding remained the same. In addition, Propedia was able to detect similar contacts in the protein–peptide interactions, suggesting conservation in the mechanism of recognition. Additionally, our analysis showed that the protein residues were conserved, but the residues of the peptides were not. However, the interaction patterns were maintained.
This study case highlights the potential of Propedia to find similar binding patterns between proteins and peptides, even when peptide primary structure is not conserved.
SARS-CoV-2 main protease interactions with peptides (6LU7)
From a SARS-CoV-2 main protease structure (PDB id: 6lu7) we performed a case study to find peptides that can potentially recognize the binding site and inhibit it competitively. We submitted the 6lu7 structure to the Propedia database webserver using CCD searching scope, setting the chain (A) and binding site residues (residues within 6 Å of the N3 inhibitor). We searched for complexes with similar binding sites and the best results were ranked by alignment score (result value from the ProBiS). The top 10 results were retrieved.
From these 10 peptides retrieved in complex with similar binding sites, we obtained the peptides presented in Table 4. We verified that both of 1lvb peptides (chains C and D) have the same sequence and structural conformation. Therefore, only the complex with 1lvb-D peptide was kept for the next analyses, which has a better bound receptor (chain B), based on it is alingment score and RMSD from Propedia query results.
Propedia was able to retrieve 2 proteases from SARS-CoV (previous coronavirus infecting human beings) and other viral proteases along with peptides that could be useful for the design of antiviral peptides capable of inhibiting the SARS-CoV-2 main protease. Consequently, we performed molecular docking experiments with the peptides returned by the search on SARS-CoV-2 protease. We used the Rosetta FlexPepDock docking protocol [44]. It computes high-resolution complex structures from an approximate model of a peptide within a receptor binding site, allowing full flexibility of the peptide backbone and all side chains. To provide the initial structure of each complex, we superposed the SARS-CoV-2 protease (PDB id: 6lu7) with the Propedia retrieved complex and removed the protein retaining SARS-CoV-2 and the peptide. This procedure was successful for 8 complexes, in which the SARS-CoV-2 protease was properly superposed with the model receptors (manual inspection and \(RMSD \le 3\) Å). For two peptides whose receptors did not align properly due to structural high dissimilarity (PDB id: 3caa:B and 6hgj:B), we performed a global blind docking using HADDOCK [45]. Then, we selected the best model from the best cluster (most negative HADDOCK score) and submitted it as thanbe initial structure to Rosetta FlexPepDock protocol as we did with the previous 8 peptides. We had to discard both peptides (PDB id: 3caa:B and 6hgj:B) because FlexDock accommodates only peptides shorter than 30 residues. Consequently, we obtained 8 docked models, and all of them exhibited considerable affinity to the SARS-CoV-2 main protease (Table 5, column “Rosetta score”). In addition to acceptable scores, we verified apparently adequate poses (Fig. 5) of each peptide for cleavage by the site considering the proximity (\(\alpha\)-C of P1) to CYS145’s sulfur atom (Table 5, column “RosettaCYS distance”).
In fact, according to the MEROPS database of proteolytic enzymes [46, 47], SARS coronavirus main proteases show preference for substrates of the general form: P4=V/T/A/S P3=V/W/K P2=L P1=H/Q. These positions are depicted in shades of blue in Fig. 4. According to these previous works, the site P1 is well conserved but the other sites are very mutable. The peptides identified using Propedia have residues highlighted in yellow and it can be viewed in the webserver (https://bioinfo.dcc.ufmg.br/propedia/search/binding/covid). Notice that this set of peptides is generally consistent with peptides known to inhibit SARS coronavirus proteases.
Metadynamics estimated \(\Delta \hbox {G}_{{bind}}\) correlates with the major propedia scores for the Sars-Cov-2 \(\hbox {M}_{{Pro}}\)
The free energy landscape (FEL) for the respective triplicates of the unbinding metadynamics (MetaD) for the \(\hbox {M}_{{Pro}}\): peptides complexes with the PDB id: 2q6g (chain C), 1uk4 (chain H), 1lvm (chain D), and 1lvb (chain D) are shown on Additional file 1: Figure S.1 (maps 1–3, 4–6, 7–9 and 10–12, respectively). At each system, the minima inside the protein (A) and at the aqueous environment (B) could be characterized with enough accuracy in order to estimate the binding free energy (\(\Delta \hbox {G}_{{bind}}\)) according the described on equations (S.1, S.2, and S.3) from the Additional file 1.
It could be obtained a significant convergence for the MetaD recovered \(\Delta \hbox {G}_{{bind}}\) values for each system in our protocol, with maximal standard deviation of 1.57 kcal\(\,\hbox {mol}^{-1}\) for the systems 1lvm (chain D), 1lvb (chain D) and 2q6g (chain C) and a relatively higher deviation of 4.32 kcal\(\,\hbox {mol}^{-1}\) just for 1uk4. In fact, such convergence is not surprising, once the already consolidated situation of the metadynamics technique as an accurate computational tool to estimate the binding free energy for usual ligands and peptides, being a powerful method on drug screening procedures [48,49,50]. The accuracy of this technique, in this way, makes it a providential instrument to validate the Propedia methodology at the screening of peptides with differential affinities for the Sars-Cov-2 MPro, given the still sparse availability of experimental data for peptide affinity at this new and important target. In this way, the correlation between the \(\Delta \hbox {G}_{{bind}}\) recovered by the metadynamics higher performance method and the Propedia recovered scores was carried aiming the validation of this computational tool. It is notorious at Fig. 6 (Additional file 1: Table S.1), the significant negative correlation of the MetaD \(\Delta \hbox {G}_{{bind}}\) with the Propedia recovered alignment score (\(\hbox {R}^2\) of 0.98) and the positive correlation with the Propedia recovered RMSD in Å at the active site alignment procedure (\(\hbox {R}^2\) of 0.96). Even, it is noteworthy that both the Propedia scores as the MetaD recovered \(\Delta \hbox {G}_{{bind}}\) values put the known MPro specific substrate (PDB:2q6g) and the substrate-analogous MPro inhibitor (1uk4) at the top of the affinity ranking with this protein. In this way, both the significant correlation with the results from the high performance metadynamics method, as well the self-consistence with known functional data can be taken together as an indicative of validation for our new software, as well its applicability at the screening for functional peptides for this and other important targets.
Anticarsia gemmatalis protease
The velvetbean caterpillar, Anticarsia gemmatalis (AG) Hübner (Lepidoptera: Noctuidae) is one of the primary defoliating pests in the Americas, affecting mainly soybean crops, and a major cause of economic losses in agriculture [51, 52]. In recent years, alternative approaches towards pest control, such as the development of biopesticides, have been explored. For instance, the use of protease inhibtors is high regarded in insect pest management, as it affects the bioavailability of essential amino acids, which ultimately hinders larvae growth and the development of insects for several species, as has been shown in [53, 54].
In this case study, we used the sequence of a trypsin-like serine protease extracted from the AG’s gut, sequenced by our research group and deposited at GenBank [55] (accession JX898746.1 [56]). Additionally, a 3D model was produced using the I-TASSER server [57]. We performed a structural alignment of the model with the highest ranked templates from the modeling step in order to identify the highly conserved residues from the catalytic triad in the protease [58]. These residues were identified as HIS6, ASP56, and SER143 in the model. Additional file 1: Figure S.2 shows the superposition of the structures where the triad residues are highlighted.
We queried the Propedia database server using the protease sequence and the residues from the catalytic triad as binding site residues (along with the 3D model) in two separate experiments, and the top 10 results in each of them were selected according to their alignment scores. The results are shown in Tables 6 and 7 respectively. Then, we performed molecular docking experiments of the peptides retrieved to the model of the AG’s protease using only HADDOCK, since a considerable number of the peptides retrieved contained non-standard residues, which is not supported by PepFlexDock. Also, for the binding site query experiment, two peptides entries from the propedia results were not used (which are not listed in Table 7): 1p11-P, due to its format not being supported by HADDOCK and 3kf2-D, due to the high sequence similarity to peptide 3kf2-C in the same PDB structure.
For the sequence based dataset, we set the residues from the catalytic triad as active residues for the docking procedure, as well as the complete chain of each peptide. We selected the best resulting structures primarily according to the HADDOCK score (most negative) and then, according to the RMSD (\(\le 3\) angstroms) of each structure relative to the overall lowest energy model. Table 8 summarize the results. Finally, peptide poses in the protease were analysed for the top 5 scored models according to the HADDOCK score, for which we identified the closest residues to the SER143 residue at the S1 site, considering the distance between C-\(\alpha\) atoms. The closest residues found were cysteine residues located in models 3qgn-A (3.9 Å) , 4dii-L (4.4 Å) and 1ca8-A (5.1 Å). The presence of cysteine residues close to the serine in the catalytic tryad indicate a potential use of the peptide as an inhibitor since substrates with these residues at position P1 are not usually cleaved by trypsin-like serine proteases [59]. Figure 7 shows models 3qgn-A and 4dii-L, where the distance between residues is highlighted.
Similar to the sequence based dataset, we performed the docking for the binding site dataset using the residues from the catalytic triad, as well as complete peptide chains as active residues. A binding site signature is how a protein interacts with its ligand, and which amino acids are essential to keep the complex stable. A proper metric to verify the similarity of binding sites is the fraction of common contacts (FCC). The \(FCC_{AB}\) is the ratio of contacts between structures A and B to all contacts in A, whose value ranges from zero, when the chains share no contacts, to a maximum of one, when all contacts of chain A are with chain B [60]. Therefore, for the binding site docking experiments, a higher average value of FCC in a cluster indicates higher similarity of the interactions between different peptide poses and the protease model, which also means that the binding site is more conserved.
For each peptide, we selected the cluster with the highest FCC score (relative to its lowest energy models produced by HADDOCK), from which sets we chose the best models according to their HADDOCK scores. FCC values and HADDOCK scores are shown in Table 9 for all peptides. The best 4 models for each of the top 3 scored clusters are shown in Fig. 8. In all models, contacts are centered in the catalytic triad (highlighted in red), while the remaining contact areas bind to different ligands, where neighboring residues on the protease side have great relevance by establishing hydrophobic and hydrogen bonds. The complete interaction map of each model is available in Additional file 1: Figure S.3. This emphasizes the importance of using FCC as a suitable metric for binding site analysis rather than RMSD, and also demonstrates Propedia’s accuracy in determining binding site patterns in regard to the ligand specificity.
Conclusions
As far as we know, Propedia is the broadest and most comprehensive set of protein–peptide complexes. At the moment of publication of this paper, it comprises approximately 20,000 complexes. Furthermore, we developed hybrid clustering strategies that organized data into 1845 clusters based on sequences, 1891 clusters based on interface structures similarity and 1466 clusters based on binding sites. These groups may be used for detecting either nonredundant or similar complexes with several purposes going from peptide docking and scoring function benchmarking, design of biotechnological peptides and even peptide-based rational drug design. Finally, Propedia is available through a web interface, searches and analysis can be performed by a user-friendly interface and all the data are available to download.
Availability of data and materials
The data generated and/or analyzed for current study are available at Propedia’s download page: https://bioinfo.dcc.ufmg.br/propedia/download
Abbreviations
- ASA::
-
Accessible surface area
- AG::
-
Anticarsia gemmatalis
- CCD::
-
Clusterable Complex Dataset
- FCC::
-
Fraction of common contacts
- FEL::
-
Free energy landscape
- IA::
-
Interface area
- MetaD::
-
Metadynamics
- NMR::
-
Nuclear magnetic resonance
- PDB::
-
Protein Data Bank
- RMSD::
-
Root mean square deviation
- iRMSD::
-
Interface root mean square deviation
References
Neduva V, Linding R, Su-Angrand I, Stark A, De Masi F, Gibson TJ, Lewis J, Serrano L, Russell RB. Systematic discovery of new recognition peptides mediating protein interaction networks. PLoS Biol. 2005;3(12):e405.
Liu D, Angelova A, Liu J, Garamus VM, Angelov B, Zhang X, Li Y, Feger G, Li N, Zou A. Self-assembly of mitochondria-specific peptide amphiphiles amplifying lung cancer cell death through targeting the vdac1-hexokinase-ii complex. J Mater Chem B. 2019;7(30):4706–16.
Lau JL, Dunn MK. Therapeutic peptides: historical perspectives, current development trends, and future directions. Bioorganic Med Chem. 2018;26(10):2700–7.
Angelova A, Drechsler M, Garamus VM, Angelov B. Pep-lipid cubosomes and vesicles compartmentalized by micelles from self-assembly of multiple neuroprotective building blocks including a large peptide hormone pacap-dha. ChemNanoMat. 2019;5(11):1381–9.
Lee AC-L, Harris JL, Khanna KK, Hong J-H. A comprehensive review on current advances in peptide drug development and design. Int J Mol Sci. 2019;20(10):2383.
Vinogradov AA, Yin Y, Suga H. Macrocyclic peptides as drug candidates: recent progress and remaining challenges. J Am Chem Soc. 2019;141(10):4167–81.
Pant S, Singh M, Ravichandiran V, Murty U, Srivastava HK. Peptide-like and small-molecule inhibitors against Covid-19. J Biomol Struct Dyn (just-accepted), 1–15 (2020)
Wang J, Yin T, Xiao X, He D, Xue Z, Jiang X, Wang Y. Strapep: a structure database of bioactive peptides. Database. 2018;2018.
Wang G, Li X, Wang Z. Apd3: the antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 2016;44(D1):1087–93.
Gautam A, Singh H, Tyagi A, Chaudhary K, Kumar R, Kapoor P, Raghava G. Cppsite: a curated database of cell penetrating peptides. Database. 2012;2012.
Gautam A, Chaudhary K, Singh S, Joshi A, Anand P, Tuknait A, Mathur D, Varshney GC, Raghava GP. Hemolytik: a database of experimentally determined hemolytic and non-hemolytic peptides. Nucleic Acids Res. 2014;42(D1):444–9.
London N, Movshovitz-Attias D, Schueler-Furman O. The structural basis of peptide-protein binding strategies. Structure. 2010;18(2):188–99.
Vanhee P, Reumers J, Stricher F, Baeten L, Serrano L, Schymkowitz J, Rousseau F. Pepx: a structural database of non-redundant protein-peptide complexes. Nucleic Acids Res. 2010;38(suppl–1):545–51.
Das AA, Sharma OP, Kumar MS, Krishna R, Mathur PP. Pepbind: a comprehensive database and computational tool for analysis of protein–peptide interactions. Genom Proteom Bioinform. 2013;11(4):241–6.
Frappier V, Duran M, Keating AE. Pixeldb: protein–peptide complexes annotated with structural conservation of the peptide binding mode. Protein Sci. 2018;27(1):276–85.
Wen Z, He J, Tao H, Huang S-Y. Pepbdb: a comprehensive structural database of biological peptide–protein interactions. Bioinformatics. 2019;35(1):175–7.
Wallace AC, Laskowski RA, Thornton JM. Ligplot: a program to generate schematic diagrams of protein–ligand interactions. Protein Eng Des Sel. 1995;8(2):127–34.
Xu X, Zou X. Peppro: a nonredundant structure data set for benchmarking peptide-protein computational docking. J Comput Chem. 2020;41:362–9.
Pilon FM, Silva CdR, Visôtto LE, Barros RdA, da Silva Júnior NR, Campos WG, de Almeida Oliveira MG. Purification and characterization of trypsin produced by gut bacteria from anticarsia gemmatalis. Archiv Insect Biochem Physiol. 2017;96(2):21407.
Patarroyo-Vargas AM, Merino-Cabrera YB, Zanuncio JC, Rocha F, Campos WG, de Almeida O, Maria G. Kinetic characterization of anticarsia gemmatalis digestive serine-proteases and the inhibitory effect of synthetic peptides. Protein Peptide Lett. 2017;24(11):1040–7.
Queiroz FC, Vargas AM, Oliveira MG, Comarela GV, Silveira SA. ppigremlin: a graph mining based detection of conserved structural arrangements in protein–protein interfaces. BMC Bioinform. 2020;21:1–25.
Cock PJ, Antao T, Chang JT, Chapman BA, Cox CJ, Dalke A, Friedberg I, Hamelryck T, Kauff F, Wilczynski B, et al. Biopython: freely available python tools for computational molecular biology and bioinformatics. Bioinformatics. 2009;25(11):1422–3.
Hamelryck T, Manderick B. Pdb file parser and structure class implemented in python. Bioinformatics. 2003;19(17):2308–10.
Fassio AV, Santos LH, Silveira SA, Ferreira RS, de Melo-Minardi RC. napoli: a graph-based strategy to detect and visualize conserved protein–ligand interactions in large-scale. In: IEEE/ACM transactions on computational biology and bioinformatics (2019).
Lee B, Richards FM. The interpretation of protein structures: estimation of static accessibility. J Mol Biol. 1971;55(3):379–3794.
Hubbard SJ, Thornton JM. Naccess. Computer Program, Department of Biochemistry and Molecular Biology, University College London, vol. 2, no. 1 (1993).
Krejci A, Hupp TR, Lexa M, Vojtesek B, Muller P. Hammock: a hidden Markov model-based peptide clustering algorithm to identify protein-interaction consensus motifs in large datasets. Bioinformatics. 2016;32(1):9–16.
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, et al. Fast, scalable generation of high-quality protein multiple sequence alignments using clustal omega. Mol Syst Biol. 2011;7(1):539.
Sievers F, Higgins DG. Clustal omega. Curr Protoc Bioinform. 2014;48(1):3–13.
Finn RD, Clements J, Eddy SR. Hmmer web server: interactive sequence similarity searching. Nucleic Acids Res. 2011;39(suppl–2):29–37.
Söding J. Protein homology detection by hmm-hmm comparison. Bioinformatics. 2005;21(7):951–60.
Crooks GE, Hon G, Chandonia J-M, Brenner SE. Weblogo: a sequence logo generator. Genome Res. 2004;14(6):1188–90.
Konagurthu AS, Whisstock JC, Stuckey PJ, Lesk AM. Mustang: a multiple structural alignment algorithm. Proteins Struct Funct Bioinform. 2006;64(3):559–74.
Bickerton GR, Higueruelo AP, Blundell TL. Comprehensive, atomic-level characterization of structurally characterized protein–protein interactions: the piccolo database. BMC Bioinform. 2011;12(1):313.
Plaxco KW, Simons KT, Baker D. Contact order, transition state placement and the refolding rates of single domain proteins. J Mol Biol. 1998;277(4):985–94.
Hagberg A, Swart P, S Chult D. Exploring network structure, dynamics, and function using networkx. Technical report, Los Alamos National Lab. (LANL), Los Alamos, NM (United States) (2008)
Konc J, Janežič D. Probis algorithm for detection of structurally similar protein binding sites by local structural alignment. Bioinformatics. 2010;26(9):1160–8.
Schmitt S, Kuhn D, Klebe G. A new method to detect related function among proteins independent of sequence and fold homology. J Mol Biol. 2002;323(2):387–406.
Karlin S, Altschul SF. Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes. Proc Natl Acad Sci. 1990;87(6):2264–8.
Rego N, Koes D. 3dmol. js: molecular visualization with webgl. Bioinformatics. 2015;31(8):1322–4.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–10.
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. Blast+: architecture and applications. BMC Bioinform. 2009;10(1):421.
DeLano WL: PyMOL; 2002.
Raveh B, London N, Schueler-Furman O. Sub-angstrom modeling of complexes between flexible peptides and globular proteins. Proteins Struct Funct Bioinform. 2010;78(9):2029–40.
Van Zundert G, Rodrigues J, Trellet M, Schmitz C, Kastritis P, Karaca E, Melquiond A, van Dijk M, De Vries S, Bonvin A. The haddock2. 2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol. 2016;428(4):720–5.
Rawlings ND, Barrett AJ, Bateman A. Merops: the peptidase database. Nucleic Acids Res. 2010;38(suppl–1):227–33.
Goetz D, Choe Y, Hansell E, Chen Y, McDowell M, Jonsson C, Roush W, McKerrow J, Craik C. Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the sars coronavirus. Biochemistry. 2007;46(30):8744–52.
Cavalli A, Spitaleri A, Saladino G, Gervasio FL. Investigating drug-target association and dissociation mechanisms using metadynamics-based algorithms. Acc Chem Res. 2015;48(2):277–85.
Söldner CA, Horn AH, Sticht H. A metadynamics-based protocol for the determination of gpcr-ligand binding modes. Int J Mol Sci. 2019;20(8):1970.
Brandt AM, Batista PR, Souza-Silva F, Alves CR, Caffarena ER. Exploring the unbinding of l eishmania (l.) amazonensis cpb derived-epitopes from h 2 mhc class i proteins. Proteins Struct Funct Bioinform. 2016;84(4):473–87.
Vianna U, Pratissoli D, Zanuncio J, Alencar J, Zinger FD. Espécies e/ou linhagens de trichogramma spp. (hymenoptera: Trochogrammatidae) para o controle de anticarsia gemmatalis (lepidoptera: Noctuidae). Arquivos do Instituto Biológico. 2011;71:81–7.
Moscardi F, Bueno A, Sosa-gÓmez D, Roggia S, Hoffmann-campo C, Pomari A, Corso I, Yano S. Capítulo 4-artrópodes que atacam as folhas da soja. Soja-manejo integrado de insetos e outros artrópodes-praga. Brasília: Embrapa, 213–334 (2012)
Moreira L, Campos W, Ribeiro F, Guedes R, Oliveira M. Survival and developmental impairment induced by the trypsin inhibitor bis-benzamidine in the velvetbean caterpillar (anticarsia gemmatalis). Crop Prot. 2011;30(10):1285–90.
Pilon AM, Campos WG, Silva CR, Cordeiro G, Silva CR, Oliveira MGA. Protease inhibitory, insecticidal and deterrent effects of the trypsin-inhibitor benzamidine on the velvetbean caterpillar in soybean. Anais da Academia Brasileira de Ciências. 2018;90(4):3475–82.
Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Wheeler DL. Genbank. Nucleic Acids Res. 2005;33(suppl–1):34–8.
GenBank Internet, Bethesda MD. National Library of Medicine (US), National Center for Biotechnology Information;1982. https://www.ncbi.nlm.nih.gov/nuccore/JX898746.1
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The i-tasser suite: protein structure and function prediction. Nat Methods. 2015;12(1):7.
Perona JJ, Craik CS. Structural basis of substrate specificity in the serine proteases. Protein Sci. 1995;4(3):337–60.
Page MJ, Di Cera E. Serine peptidases: classification, structure and function. Cell Mol Life Sci. 2008;65(7–8):1220–36.
Rodrigues JP, Trellet M, Schmitz C, Kastritis P, Karaca E, Melquiond AS, Bonvin AM. Clustering biomolecular complexes by residue contacts similarity. Proteins Struct Funct Bioinform. 2012;80(7):1810–7.
Acknowledgements
Not applicable
Funding
This study was supported in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Grant 51/2013 - 23038.004007/2014-82; Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq); Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG). The funding body did not play any roles in the design of the study, collection, analysis, or interpretation of data or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
RCMM, SAS, MGAO and PMM conceived the study. PMM, LHS, DM and RCMM wrote the manuscript. PMM and DM developed the webtool. PMM performed sequence, interface and binding site clustering. LHS, DM, and MTQM performed the case study of estrogen receptor complexes. LLB, RCMM, LHFL, PHCF, and REOR performed docking analysis. LHFL, PHCF, and REOR performed metadynamics validation for the case study of SARS-CoV-2 main protease. FCQ and ISG performed the case study of Anticarsia gemmatalis protease. PMM, RCMM and SAS revised the manuscript. RCMM: funding acquisition. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable
Consent for publication
Not applicable
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1.
Additional details and figures for casestudies: Metadynamics estimated \(\Delta\hbox{G}_{{bind}}\) correlateswith the major propedia scores for the Sars-Cov-2 \(\hbox{M}_{{Pro}}\) and Anticarsia gemmatalis protease.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Martins, P.M., Santos, L.H., Mariano, D. et al. Propedia: a database for protein–peptide identification based on a hybrid clustering algorithm. BMC Bioinformatics 22, 1 (2021). https://doi.org/10.1186/s12859-020-03881-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12859-020-03881-z